Mechanisms of Protein Efflux
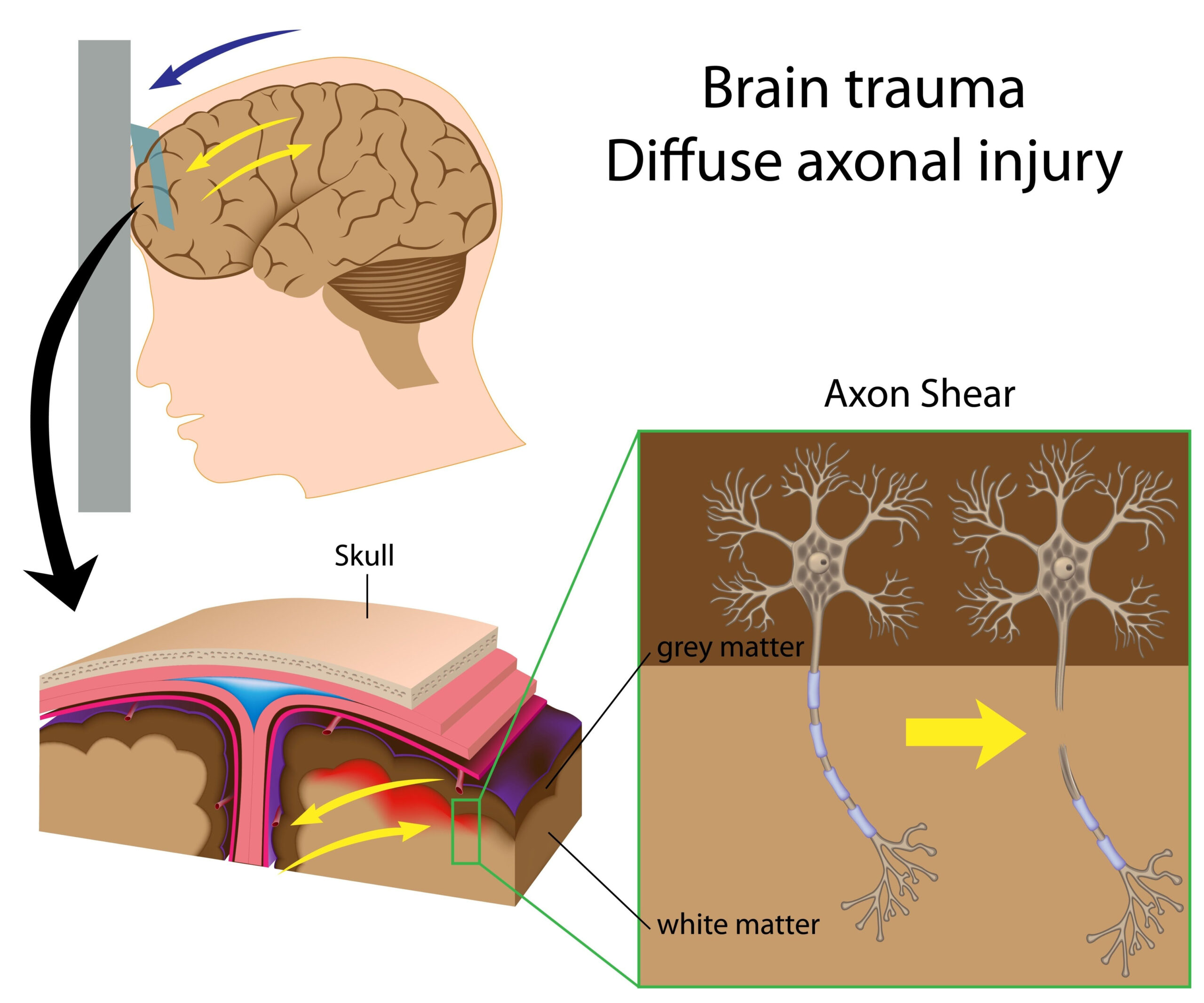
Traumatic brain injury (TBI) triggers a complex cascade of cellular responses, significantly affecting the dynamics of protein release from neurons and glial cells. After a traumatic event, cellular integrity is compromised, leading to alterations in the blood-brain barrier (BBB) and increased permeability of neuronal membranes. This permeability shift can facilitate both the efflux of distinct proteins from the intracellular environment into the extracellular space and their entry into the bloodstream.
One of the primary mechanisms involved in the efflux of proteins is vesicular transport, where proteins are packaged into vesicles that fuse with the plasma membrane and release their contents outside the cell. Key proteins, such as amyloid precursor protein (APP) and various neurotrophic factors, utilize this pathway to exit neuronal cells. This mechanism is crucial in the context of neurodegenerative processes, where the accumulation of these proteins may influence neuroinflammatory responses post-injury.
In addition to vesicular transport, transmembrane transport mechanisms also play a significant role. Channel proteins and transporters that reside in neuronal cell membranes can mediate the movement of larger proteins directly out of cells. For instance, the release of glutamate, a key neurotransmitter, can lead to excitotoxicity and further neuronal damage if not tightly regulated. Proteins may also be expelled through ATP-binding cassette (ABC) transporters, which actively efflux a variety of substrates, including signaling peptides and toxic metabolites, thus serving as a protective mechanism in the face of cellular stress.
Research also highlights the involvement of pathological processes like oxidative stress, which can alter the normal functioning of these transport mechanisms, potentially exacerbating neuronal damage and contributing to protein accumulation within the brain. Notably, elevated levels of oxidative stress can lead to lipid peroxidation, affecting membrane fluidity and potentially enhancing protein efflux.
Understanding the intricate mechanisms governing protein efflux following TBI is essential, not only for uncovering how neurons respond to injury but also for exploring therapeutic interventions aimed at modulating this process. Enhanced clarity on these mechanisms can aid in the development of strategies to either promote beneficial efflux of neuroprotective proteins or prevent the misguided release of potentially harmful proteins that could worsen injury outcomes.
Barriers to Understanding
The investigation of protein efflux in the context of traumatic brain injury (TBI) encounters several significant barriers that impede holistic understanding and effective therapeutic advancements. One primary barrier is the complexity of the brain’s microenvironment, which comprises various cell types, including neurons, astrocytes, and microglia, all interacting dynamically in response to injury. These intricate relationships complicate the isolation and identification of specific proteins involved in the efflux process. The interplay of multiple signaling pathways further obscures clear cause-and-effect relationships, making it challenging to discern which proteins are most critical to outcomes following TBI.
Additionally, the physiological state of the blood-brain barrier (BBB) itself presents an obstacle. Following TBI, the BBB often becomes disrupted, leading to increased permeability which can result in indiscriminate protein efflux. However, this damage varies greatly between individuals and even across different regions of the brain within the same individual. Such variability can skew research results and complicate the standardization of experimental models used to study these processes. Differences in the timing of injury, the severity of the trauma, and the resultant cellular responses can yield inconsistent data, hampering our ability to generalize findings across populations.
Another significant barrier is the current methodologies employed in research. Traditional techniques for analyzing protein levels, such as ELISA (enzyme-linked immunosorbent assay) or Western blotting, often lack the sensitivity required for detecting low abundance proteins in the context of neurotrauma. Moreover, these methods typically analyze homogenized brain tissue rather than specific cell types, obscuring the contributions of individual cellular responses. Advances in single-cell transcriptomics and proteomics offer promise, yet these technologies are still developing and may not be widely accessible.
Furthermore, the timeline of protein efflux relative to the timing of TBI is a critical detractor from our understanding. Protein release can occur acutely following injury or may be a delayed response that continues for days or even weeks post-trauma. This temporal aspect complicates the study of the proteins involved, as researchers may fail to capture transient events that are crucial to understanding the overall efflux dynamics.
Ethical considerations also arise when designing studies involving human subjects, particularly those with acute TBI. Obtaining timely biopsies or cerebrospinal fluid samples is often impractical and can hinder the ability to assess protein levels effectively during critical post-injury windows. Additionally, the reliance on animal models, while necessary for certain experimental designs, introduces another layer of complexity given the differences between species in neurobiology and the potential limitations in translating findings to human populations.
In summary, the multifaceted nature of both the biological environment and research methodologies presents significant challenges to our understanding of protein efflux following TBI. Overcoming these barriers is essential for advancing our knowledge and developing targeted therapeutic strategies that could mitigate the long-term consequences of traumatic brain injuries. Addressing these hurdles requires interdisciplinary collaboration within the scientific community to refine techniques, enhance model systems, and incorporate a range of biological complexities into future research efforts.
Experimental Approaches
Exploration of protein efflux mechanisms following traumatic brain injury (TBI) necessitates a diverse array of experimental methodologies that can adequately capture the dynamic processes occurring within the injured brain. A multifaceted approach is critical, as it enables researchers to gather comprehensive data that elucidates the pathways through which proteins are released into the extracellular environment, thereby facilitating a clearer understanding of post-injury neuronal responses.
One of the foundational methods employed in studying protein dynamics is immunohistochemistry, which allows for the visualization of specific proteins within brain tissue sections. Through the use of labeled antibodies, researchers can observe the spatial expression of proteins that may be involved in the efflux process, such as APP and neurotrophic factors. This technique provides insights into the localization and potential release sites of these proteins, although it is limited in quantifying protein levels and detecting low-abundance proteins.
In addition to immunohistochemistry, mass spectrometry offers a powerful tool for identifying and quantifying proteins in complex biological samples. By analyzing the proteome of brain tissues collected post-TBI, researchers can identify the specific proteins that are differentially expressed following injury. Mass spectrometry can facilitate the detection of both abundant and low-abundance proteins, thereby shedding light on previously unnoticed pathways of protein efflux. Moreover, this technique can be further enhanced through the use of liquid chromatography tandem mass spectrometry (LC-MS/MS), allowing for deeper profiling of protein modifications and interactions.
To complement these biochemical analyses, the use of in vivo and in vitro models of TBI provides a platform for studying protein efflux in real-time under controlled conditions. Animal models, such as rodents subjected to controlled cortical impact or blast injury, enable researchers to monitor the temporal dynamics of protein release immediately following injury. In vitro systems utilizing cultured primary neurons or glial cells exposed to simulated traumatic conditions present researchers the opportunity to dissect cellular mechanisms and identify key signaling pathways that regulate protein efflux. Additionally, the development of organoid models derived from human neural tissue offers an innovative avenue to study the complexities of protein dynamics in a more physiologically relevant context.
Comparative studies utilizing both acute and chronic models of TBI can yield insights into the temporal aspects of protein efflux, allowing scientists to discern patterns in protein release that may correlate with neuronal recovery or degeneration. This longitudinal approach enables researchers to collect data on protein levels at various stages post-injury, providing a rich dataset for understanding the relationship between effluxed proteins and functional outcomes.
Importantly, the advent of advanced imaging techniques, such as two-photon microscopy and live-cell imaging, facilitates real-time observation of protein transport dynamics within the brain. These methods can reveal intricate details on how proteins are secreted from cells and their subsequent interactions with the surrounding environment. Such capabilities are crucial for exploring the immediate effects of TBI on protein release and cellular behavior.
Finally, collaborations across disciplines—from molecular biology to bioengineering—are essential for developing novel methods to overcome current research limitations. Implementing gene editing technologies, such as CRISPR/Cas9, for targeted manipulation of genes involved in protein transport can illuminate the specific contributions of different proteins in the context of TBI. Additionally, high-throughput screening methods could allow for the testing of numerous compounds that might modulate protein efflux, paving the way for potential therapeutic discoveries.
Collectively, these experimental approaches not only enhance our understanding of the mechanisms governing protein efflux following TBI but also open avenues for identifying targets for therapeutic intervention. By integrating these diverse methodologies, researchers can build a comprehensive understanding that ultimately informs strategies to mitigate the long-term consequences of brain injury.
Future Directions
As research into the protein efflux mechanisms following traumatic brain injury (TBI) progresses, several promising avenues warrant further exploration to enhance our understanding and develop potential therapeutic strategies. Identifying novel biomarkers that can be reliably measured in cerebrospinal fluid (CSF) or blood could provide critical insights into the timing and extent of protein efflux, offering a non-invasive means to monitor neurotrauma and recovery. This biomarker approach could lead to improved diagnostic tools and facilitate timely intervention strategies in clinical settings.
Advancements in technology will also play a pivotal role in driving future research. The continued refinement of single-cell sequencing techniques will allow researchers to dissect the cellular responses of specific neuronal and glial populations following TBI, providing a more granular view of how different cell types contribute to protein dynamics. Coupled with sophisticated bioinformatics tools, these approaches will enable better integration of large datasets, elucidating complex signaling networks involved in protein trafficking.
In addition, the development of targeted therapeutics that can either enhance beneficial protein efflux or inhibit harmful protein release presents an exciting direction. Research on small molecule modulators or monoclonal antibodies that can specifically influence transport mechanisms may pave the way for interventions that protect neuronal integrity after an injury. For example, exploring the manipulation of ABC transporters or vesicular pathways could yield strategies aimed at fostering neuroprotection while minimizing the risk of exacerbating injury.
The exploration of exosomes and microvesicles as vehicles for protein efflux is another intriguing pathway. Understanding how these extracellular vesicles are involved in the transport of biomolecules following TBI could lead to the identification of novel therapeutic targets. Further studies on the role of these vesicles in cell-to-cell communication in the injured brain may reveal how the efflux of certain proteins influences nearby cells and overall brain recovery.
Another promising area of research involves the exploration of the neuroinflammatory response post-TBI. Understanding how activated glial cells influence protein efflux could provide insights into the balance between neuroprotection and neurodegeneration. Investigating the nature of the inflammatory milieu and how it interacts with neuronal cells to modulate protein release may unlock new therapeutic avenues aimed at tuning the inflammatory response following injury.
Clinical translation of these findings remains a critical component of future research. Collaborative efforts between basic scientists, clinicians, and engineers will be essential in bridging the gap from laboratory discoveries to standardized clinical applications. Initiatives that focus on building robust preclinical and clinical trials will be vital to translating the knowledge gained about protein efflux mechanisms into therapies that truly benefit TBI patients.
As we look to the future, interdisciplinary collaboration, technological innovation, and a focus on patient-centered research will be key to overcoming existing barriers. By addressing the complex interplay between protein dynamics, cellular responses, and therapeutic interventions, we can move closer to improving outcomes for individuals affected by traumatic brain injuries.