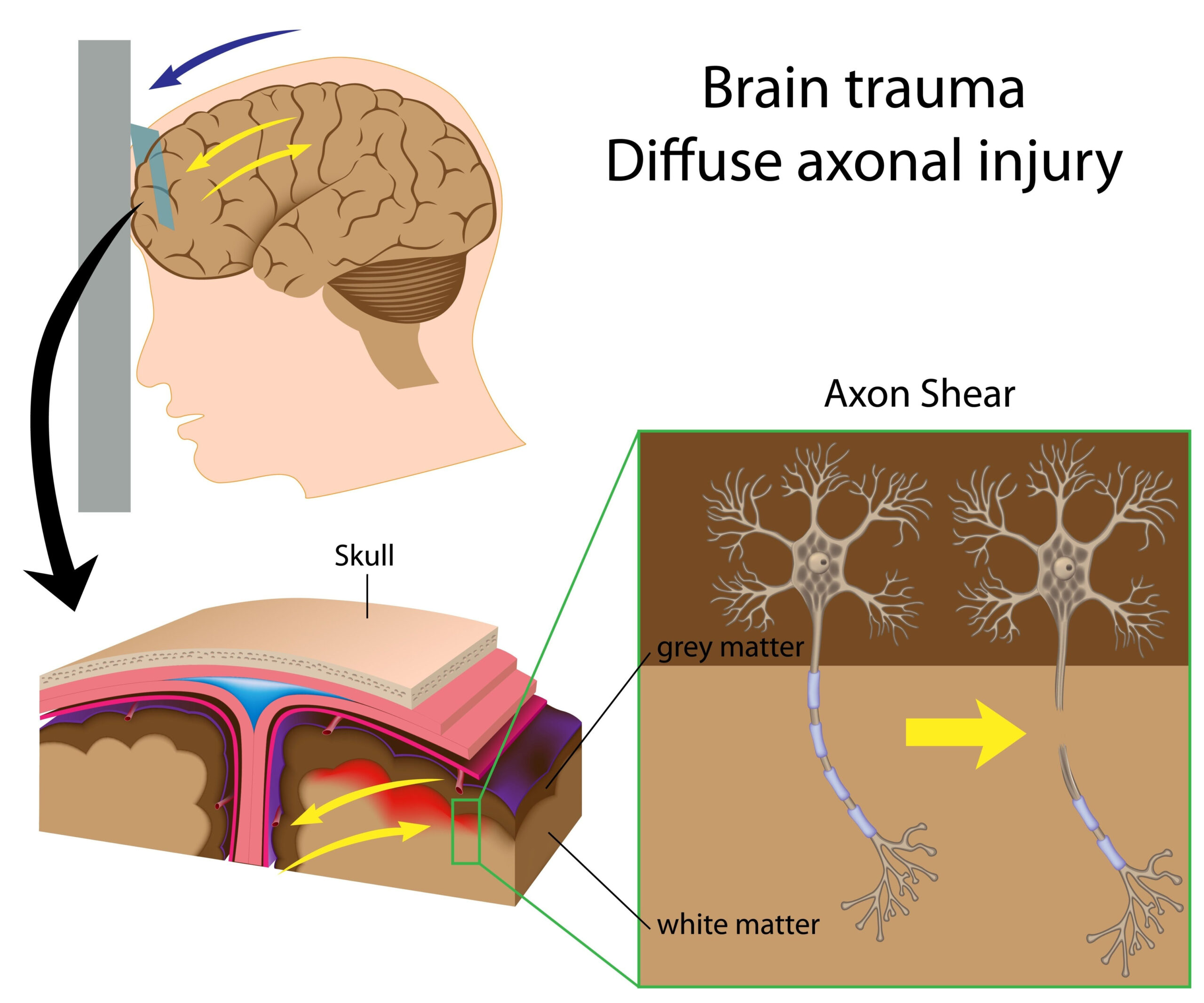
Transcriptomic Changes Induced by Injury
Mild traumatic brain injury (mTBI) triggers significant changes in the brain’s transcriptome, the complete set of RNA molecules, reflecting the genetic activity within cells. These transcriptomic alterations facilitate the brain’s response to injury and can provide insights into the underlying mechanisms of recovery and dysfunction. Following mTBI, various cell types exhibit distinct alterations in their gene expression profiles, which may be influenced by the severity and nature of the injury.
Studies using advanced sequencing technologies have uncovered widespread shifts in RNA expression post-injury. For instance, certain immune-related genes become upregulated, reflecting an activation of inflammatory pathways that are typically engaged during the healing process. Concurrently, genes associated with cellular stress response and repair mechanisms are also activated, illustrating the brain’s attempt to restore homeostasis after the injury.
Moreover, specific neuronal populations, such as excitatory and inhibitory neurons, display different patterns of transcriptomic reorganization. This differential gene expression is crucial because it not only underscores the complexity of the brain’s response but also indicates that different cell types may have specialized roles in recovery or pathogenesis following the injury. For example, in excitatory neurons, many genes involved in synaptic plasticity show altered expression, which can impact cognitive function if the reorganization is maladaptive.
Additionally, glial cells, which support and protect neuronal function, also undergo significant changes in gene expression. Astrogliosis, or the proliferation of astrocytes, is one response observed post-mTBI, driven by increased expression of certain signaling molecules and growth factors. This response can help in the repair of the blood-brain barrier and facilitate neuronal survival, although excessive glial activation can have detrimental effects.
The alterations in gene expression are not uniform; they can vary according to the time post-injury, the particular brain region affected, and the cellular context. For instance, some genes may be rapidly upregulated in response to initial injury but subsequently downregulated, highlighting the dynamic nature of the transcriptomic landscape following an mTBI.
Understanding these transcriptomic changes is essential, as they may reveal targets for therapeutic intervention aimed at enhancing recovery and minimizing long-term detrimental effects. Furthermore, these insights can guide the development of biomarkers to better assess the risk of ongoing neurological issues following mild traumatic brain injury.
Experimental Design and Techniques
The investigation of transcriptomic changes following mild traumatic brain injury (mTBI) requires a robust experimental framework that integrates various techniques to ensure accurate and comprehensive results. Typically, these studies begin with well-defined animal models, often utilizing rodents, to mimic mTBI conditions in a controlled environment. By applying a standardized impact protocol, researchers can induce injuries that reflect the characteristics of mTBI seen in humans, allowing for the observation of resultant biological processes.
One of the essential methodologies employed is RNA sequencing (RNA-Seq), which enables the researchers to capture the breadth of transcriptomic alterations by quantifying gene expression levels across different cells and time points post-injury. This high-throughput technique not only detects the abundance of mRNA but also provides information on alternative splicing and post-transcriptional modifications, giving a comprehensive view of cellular responses to injury.
To complement RNA-Seq, researchers often use real-time quantitative polymerase chain reaction (qPCR) to validate findings from sequencing data. This technique allows for the amplification and quantification of specific RNA targets that are of interest, ensuring that observed changes are statistically robust and reproducible. By choosing key genes identified during RNA-Seq analysis, researchers can focus on the most crucial aspects of the injury response.
Furthermore, immunohistochemistry and in situ hybridization techniques are employed to visualize the spatial distribution of gene expression changes within the brain. These methods enable the detection of specific proteins or RNA transcripts within tissue sections, permitting researchers to ascertain not only which genes are activated but also the specific cell types involved in the injury response. This spatial resolution is vital for understanding the localized effects of mTBI on different brain regions, as responses may vary significantly across areas such as the cortex, hippocampus, and cerebellum.
To assess functional outcomes alongside transcriptomic analysis, behavioral assays are routinely integrated into the experimental design. These tests evaluate cognitive, motor, and emotional functions, providing insights into how transcriptomic changes translate to observable alterations in behavior post-injury. Assessing these parameters allows for a more holistic understanding of the impact of mTBI on the organism, linking molecular changes to functional propensities.
Additionally, ethical considerations are paramount in studies involving animal models. All procedures must undergo institutional review board approval to ensure humane treatment of the animals. Beyond ethical practices, the reproducibility of results is essential; thus, researchers are encouraged to adhere to standardized protocols and report their methodologies transparently.
The synthesis of these techniques not only drives the progression of our understanding of mTBI but also highlights potential therapeutic targets for intervention. By elucidating the intricate transcriptomic reorganization following such injuries, researchers aim to pave the way for innovative approaches to treatment and recovery, ultimately enhancing outcomes for individuals affected by mTBI.
Post-Injury Gene Expression Patterns
Following mild traumatic brain injury (mTBI), the gene expression landscape undergoes complex reorganization that offers crucial insights into the biological processes at play during recovery. Research has shown that specific patterns of gene expression can vary significantly based on the timing of the assessment, the brain region examined, and the cell type involved, emphasizing the intricate dynamics of post-injury responses.
Early after an mTBI, certain gene categories, particularly those involved in inflammatory responses, are prominently activated. These inflammatory genes include cytokines and chemokines, which help recruit immune cells to the injury site, purportedly to facilitate repair and combat potential infection. This early inflammatory response is essential, as it activates a cascade of events aimed at restoring cellular integrity. However, if this response is excessive or poorly regulated, it can lead to secondary damage and chronic neuroinflammation, indicating a delicate balance between beneficial and harmful outcomes.
Neuronal populations demonstrate distinct shifts in gene expression that correlate with their roles in synaptic plasticity and function. For instance, genes associated with the synaptic transmission of excitatory neurotransmitters like glutamate show notable changes, which can impact cognitive functions such as learning and memory. Concurrently, inhibitory neurons can also exhibit altered expression patterns, potentially leading to imbalances in excitatory-inhibitory signaling. These imbalances may manifest as cognitive deficits or mood disorders, illustrating the critical need to monitor these gene expression changes carefully.
A significant aspect of the post-injury transcriptomic response is the activity of glial cells, particularly astrocytes and microglia. Astrogliosis is often observed, characterized by the upregulation of astrocytic genes that promote cellular proliferation and the secretion of neurotrophic factors. These changes are generally protective, aimed at restoring homeostasis and supporting neuronal survival. However, the transition to a reactive state can also be associated with harmful effects, including scarring and hindered regeneration if prolonged. This duality underscores the complexity of glial responses following injury.
Chromatin remodeling is another avenue through which gene expression profiles are altered post-mTBI. The accessibility of DNA regions for transcription can be modified, allowing for the rapid activation or silencing of genes crucial for the cell’s adaptive response following injury. Changes in epigenetic regulation mechanisms, such as DNA methylation and histone modifications, can influence which genes are expressed and to what degree, thereby shaping the cellular response to injury.
The specific timeline of gene expression changes also plays a pivotal role in the recovery trajectory. For example, certain genes may peak in expression shortly after the injury, while others may continue to rise or fall in subsequent days or weeks. Such temporal dynamics are critical for understanding the phases of recovery and identifying potential therapeutic windows where intervention may be most beneficial.
Collectively, the post-injury gene expression patterns reveal a sophisticated interplay among various cellular processes in the brain after mTBI. These insights not only enhance our understanding of the acute and subacute phases of mTBI recovery but also illuminate potential biomarkers for monitoring recovery progress and targeting therapeutic strategies effectively. By delving deeper into these patterns, researchers can better elucidate the complexities of brain injuries and their long-term impacts on neurological health.
Future Research Directions
The exploration of transcriptomic alterations following mild traumatic brain injury (mTBI) opens numerous avenues for future research that could significantly enhance our understanding of brain injuries and their consequences. An essential direction involves the longitudinal study of gene expression changes to establish a comprehensive timeline of the molecular responses to mTBI. By monitoring transcriptomic shifts across various time points, researchers could delineate acute, subacute, and chronic phases of recovery, potentially identifying critical windows for therapeutic interventions.
Moreover, expanded research into the specific roles of different cell types in the injury response will provide valuable insights. While current studies have begun to reveal unique transcriptomic profiles for various neuronal and glial populations, a more granular approach is necessary. Employing single-cell RNA sequencing technologies could allow for a detailed exploration of cellular heterogeneity, uncovering how each cell type contributes to both the repair mechanisms and the potential for maladaptive responses following injury.
Another promising direction lies in understanding the interplay between genetic and environmental factors in shaping post-injury outcomes. Individual genetic predispositions may influence how the brain responds to mTBI, impacting recovery trajectories and vulnerability to chronic conditions. Investigating gene-environment interactions could lead to personalized approaches in treatment, tailoring therapeutic strategies to meet the unique needs of each patient.
Furthermore, the integration of transcriptomic data with advanced imaging techniques presents an exciting frontier. By correlating changes in gene expression with structural and functional neuroimaging data, researchers could better understand how molecular alterations manifest in physiological and behavioral outcomes. This integrative approach could pave the way for more precise prognostic tools and therapeutic interventions following mTBI.
Targeted therapeutic strategies that emerge from transcriptomic research will also be critical. There is a need to identify specific genes or pathways that could serve as potential targets for pharmacological intervention. For instance, modulating inflammatory pathways or promoting neuroprotective gene expression through drug development may enhance recovery and mitigate chronic symptoms. Preclinical studies focusing on these targeted therapies will be vital to evaluate their efficacy and safety before human application.
Additionally, exploring the role of lifestyle and rehabilitation interventions in modulating gene expression after mTBI could enrich our understanding. Physical activity, cognitive rehabilitation, and dietary modifications have shown promise in enhancing recovery and may interact with molecular pathways to influence transcriptomic profiles. Investigating these relationships could lead to comprehensive rehabilitation strategies that harness both biological and behavioral modalities for optimal recovery.
Lastly, addressing the ethical considerations surrounding research in this area cannot be overstated. As our understanding of mTBI and its effects deepens, ensuring that research protocols respect patient autonomy and prioritize participant welfare will be essential. Moreover, fostering collaborations among scientists, clinicians, and ethicists can help navigate the complexities of conducting mTBI research in human subjects.
Future research directed at refining our understanding of transcriptomic alterations following mTBI holds the promise of revealing new therapeutic targets, improving recovery outcomes, and enhancing quality of care for individuals affected by such injuries. By embracing a multidisciplinary approach that factors in genetic, environmental, and therapeutic elements, the scientific community can advance knowledge and interventions related to mTBI.